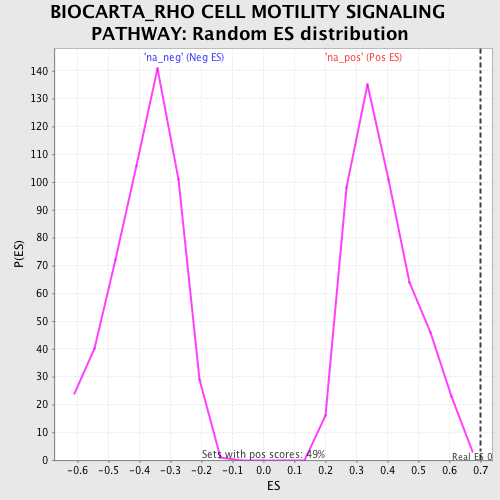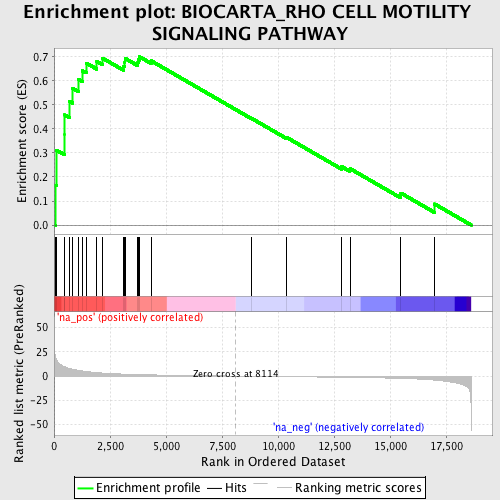
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | BIOCARTA_RHO CELL MOTILITY SIGNALING PATHWAY |
| Enrichment Score (ES) | 0.6997542 |
| Normalized Enrichment Score (NES) | 1.825826 |
| Nominal p-value | 0.002057613 |
| FDR q-value | 0.07116044 |
| FWER p-Value | 0.392 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MYLK | 70 | 19.287 | 0.1649 | Yes | ||
| 2 | ARHGEF5 | 101 | 16.999 | 0.3119 | Yes | ||
| 3 | ARFGAP3 | 476 | 9.537 | 0.3752 | Yes | ||
| 4 | VAV1 | 479 | 9.504 | 0.4582 | Yes | ||
| 5 | PIP5K1A | 694 | 7.753 | 0.5145 | Yes | ||
| 6 | RALBP1 | 832 | 7.008 | 0.5684 | Yes | ||
| 7 | NCF2 | 1099 | 5.824 | 0.6050 | Yes | ||
| 8 | PIP5K1B | 1267 | 5.221 | 0.6417 | Yes | ||
| 9 | VCL | 1435 | 4.727 | 0.6740 | Yes | ||
| 10 | GSN | 1906 | 3.624 | 0.6804 | Yes | ||
| 11 | ARHGAP4 | 2163 | 3.163 | 0.6943 | Yes | ||
| 12 | ARHGAP1 | 3104 | 2.051 | 0.6617 | Yes | ||
| 13 | MYL2 | 3157 | 2.010 | 0.6765 | Yes | ||
| 14 | ARHGEF1 | 3164 | 2.004 | 0.6937 | Yes | ||
| 15 | RHOA | 3738 | 1.602 | 0.6769 | Yes | ||
| 16 | TLN1 | 3779 | 1.568 | 0.6884 | Yes | ||
| 17 | ARHGAP6 | 3820 | 1.542 | 0.6998 | Yes | ||
| 18 | LIMK1 | 4333 | 1.259 | 0.6832 | No | ||
| 19 | ROCK1 | 8799 | -0.181 | 0.4447 | No | ||
| 20 | ARHGAP5 | 10358 | -0.571 | 0.3659 | No | ||
| 21 | CACNA1A | 12836 | -1.211 | 0.2432 | No | ||
| 22 | ARFGAP1 | 13208 | -1.323 | 0.2348 | No | ||
| 23 | PFN1 | 15475 | -2.334 | 0.1334 | No | ||
| 24 | CFL1 | 16985 | -4.061 | 0.0877 | No |